



Chemotherapy drugs lack selectivity, which means they not only kill cancer cells but also damage normal cells, leading to adverse reactions. The efficacy of chemotherapy (efficacy and toxicity of chemotherapeutic drugs ) is influenced by individual differences and the inherent toxicity of the drugs. For example, UGT1A1 affects UDP-glucuronosyltransferase/dihydropyrimidine dehydrogenase activity, which is related to the toxicity of irinotecan; variations in the DPYD gene can increase the toxicity of fluoropyrimidine drugs [1]; and the polymorphisms in XRCC1, XPD, and MTHFR genes are associated with the efficacy of platinum-based drugs in NSCLC patients [2-3].
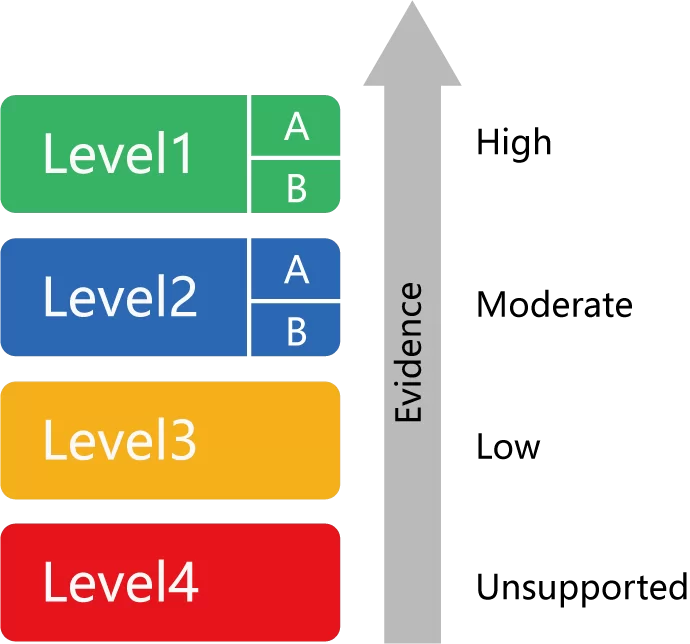
The assignment of clinical annotation levels of evidence (LOE) is primarily informed by the PharmGKB annotation scoring system for clinical annotations and variant annotations. Descriptions and scoring ranges for each level are given in the table below. Information about how PharmGKB assigns rare variant status can be found here.
1A:Level 1A clinical annotations describe variant-drug combinations that have variant-specific prescribing guidance available in a current clinical guideline annotation or an FDA-approved drug label annotation.
1B:Level 1B clinical annotations describe variant-drug combinations with a high level of evidence supporting the association but no variant-specific prescribing guidance in an annotated clinical guideline or FDA drug label.
2A:Variants in Level 2A clinical annotations are found in PharmGKB’s Tier 1 Very Important Pharmacogenes (VIPs). These variants are in known pharmacogenes, implying causation of drug phenotype is more likely.
2B:Variants in Level 2B clinical annotations are not in PharmGKB’s Tier 1 VIPs. These clinical annotations describe variant-drug combinations with a moderate level of evidence supporting the association.
3:Level 3 clinical annotations describe variant-drug combinations with a low level of evidence supporting the association.
4:Level 4 clinical annotations describe variant-drug combinations where the total score is negative and the evidence does not support an association between the variant and the drug phenotype.
[1] Chinese Clinical Oncology,Mar.2021,Vol.26,No.3
[2] Genet Mol Res. 2014 Jan 14;13(1):228-36.
[3] Pharmacogenomics. 2011 Jun;12(6):797-808.

By conducting testing on chemotherapeutic drug-related genes in cancer patients, it is possible to determine the patient's response, resistance, and the magnitude of adverse reactions to specific drugs. This scientific prediction of drug efficacy can enhance treatment targeting, avoid adverse effects, and provide valuable guidance for clinical drug therapy.
1. Nucleic Acid Extraction
2. Library Preparation (3.5 hours total time)
3. Sequencing
4. Auto-data Analysis
5. Report





 闽公网安备35021202000745
闽公网安备35021202000745