



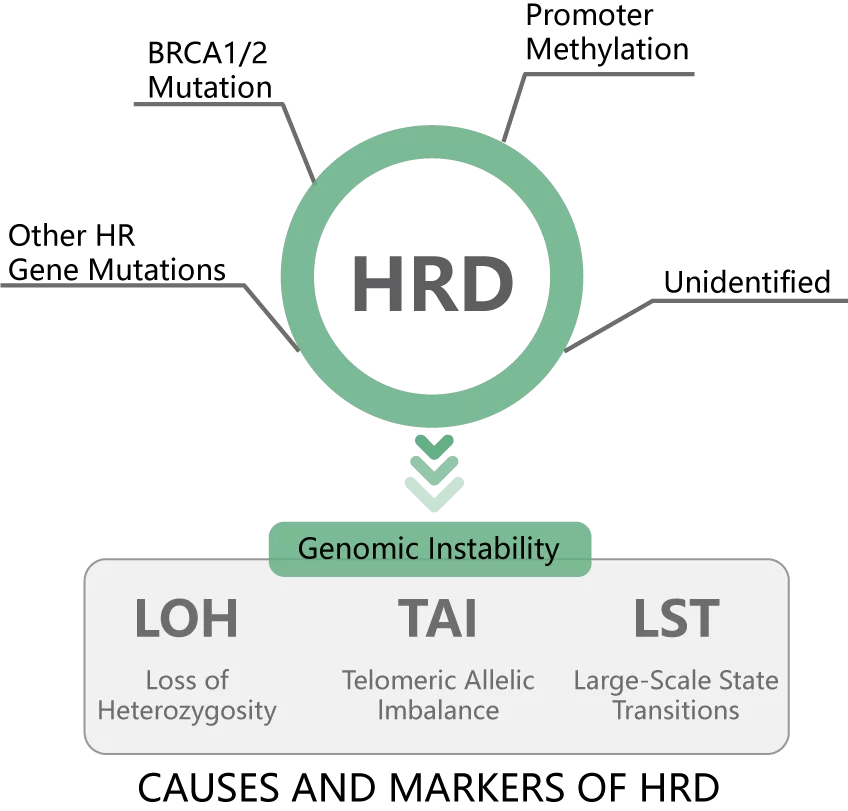
Homologous Recombination Deficiency (HRD) typically refers to a cellular-level impairment in homologous recombination repair (HRR) functionality. This condition can result from various factors, including germline mutations in HRR-related genes, somatic mutations, and epigenetic inactivation [1]. TCGA studies suggest that approximately half of high-grade serous ovarian cancer (HGSOC) may exhibit HRD, but only about 20% of patients carry pathogenic BRCA1/2 mutations. HRD can also result from other factors such as BRCA1 methylation and mutations in other HRR genes [2]. HRD can lead to specific, quantifiable, and stable genomic alterations. Loss of Heterogeneity (LOH), Telomeric Allelic Imbalance (TAI), and Large-scale State Transitions (LST), are used as indicators of Genomic Scar. The unweighted sum of these markers is used as an HRD score [3]. Combining pathogenic mutations in BRCA1/2 with the HRD score can nearly double the population benefiting from testing compared to testing for BRCA1/2 gene mutations alone.
Clinical testing for HRD has significant application value in predicting the efficacy of PARP inhibitors in the treatment of advanced ovarian cancer. It can stratify ovarian cancer patients, optimize treatment decisions, and maximize the clinical benefit of PARP inhibitors. Furthermore, in breast cancer, pancreatic cancer, and prostate cancer, HRD testing may also have potential guidance value for the clinical use of PARP inhibitors or platinum compounds [1].
FDA, EMA and NMPA have approved olaparib combined with bevacizumab for the maintenance treatment of ovarian cancer evaluated as CR / PR after first-line platinum-based chemotherapy[4]. While the indication for niraparib in the systemic treatment of heavily recurrent ovarian cancer has been withdrawn, the NCCN guidelines still maintain this recommendation [5]. Both of these treatment approaches rely on HRD as a biomarker. The NCCN Guidelines state that HRD testing is applicable to patients without gBRCA mutations to predict the efficacy of PARP inhibitors. Currently, the efficacy of PARP inhibitors in patients with Homologous Recombination Proficient (HRP) function is very limited[5].
[1] Expert consensus on clinical detection and application of homologous recombination repair defects (2021 version)
[2] Nature. 2011 Jun 29; 474(7353): 609-15.
[3] Clin Cancer Res. 2016 Aug 1;22(15):3764-73.
[4] FDA, EMA, NMPA database
[5] NCCN Ovarian Cancer, Fallopian Tube Cancer, and Primary Peritoneal Cancer Diagnosis and Treatment Guidelines 2024 v3

The proportion of tumor cells detected by tumor tissue + control samples needs to reach more than 20 %, and the proportion of tumor cells detected by single tumor tissue samples needs to reach more than 30 %.
1. Combining pathogenic mutations in BRCA1/2 with HRD score can be used to assess the HRD status of tumors, which suggests the efficacy of PARP inhibitors in patients with ovarian cancer and other patients.
2. Testing for mutations in 36 HRR genes provides comprehensive guidance on the use of PARP inhibitors in patients with prostate cancer and metastatic breast cancer.
3. Testing for mutations in 15 common driver genes offers insights into cross-cancer therapy efficacy and targeted treatment efficacy for rare ovarian cancer subtypes like low-grade serous ovarian carcinoma (LGSOC).
4.Testing for germline mutations in HRR and MMR genes can indicate the risk of hereditary cancer syndromes such as Hereditary Breast and Ovarian Cancer (HBOC) and Lynch syndrome.
HRD status may serve as a biomarker for efficacy of PARP inhibitors in the following indications:
1. Maintenance therapy for ovarian cancer evaluated as CR/PR after first-line platinum-containing chemotherapy
2. Maintenance treatment of platinum-sensitive recurrent ovarian cancer
3. Systemic treatment of multiple lines of recurrent ovarian cancer
4. Adjuvant treatment after neoadjuvant/adjuvant chemotherapy for HER2-negative early-stage high-risk breast cancer
5. Salvage treatment of HER2-negative metastatic breast cancer
6. Combined with new endocrine therapy drugs for the treatment of metastatic castration resistant prostate cancer ( mCRPC ).
7.Systemic treatment of mCRPC after failure of new endocrine therapy
8 Maintenance treatment of metastatic pancreatic cancer that has not progressed after first-line platinum-containing chemotherapy
1. Nucleic Acid Extraction
2. Library Preparation (3.5 hours total time)
3. Sequencing
4. Auto-data Analysis
5. Report





 闽公网安备35021202000745
闽公网安备35021202000745