



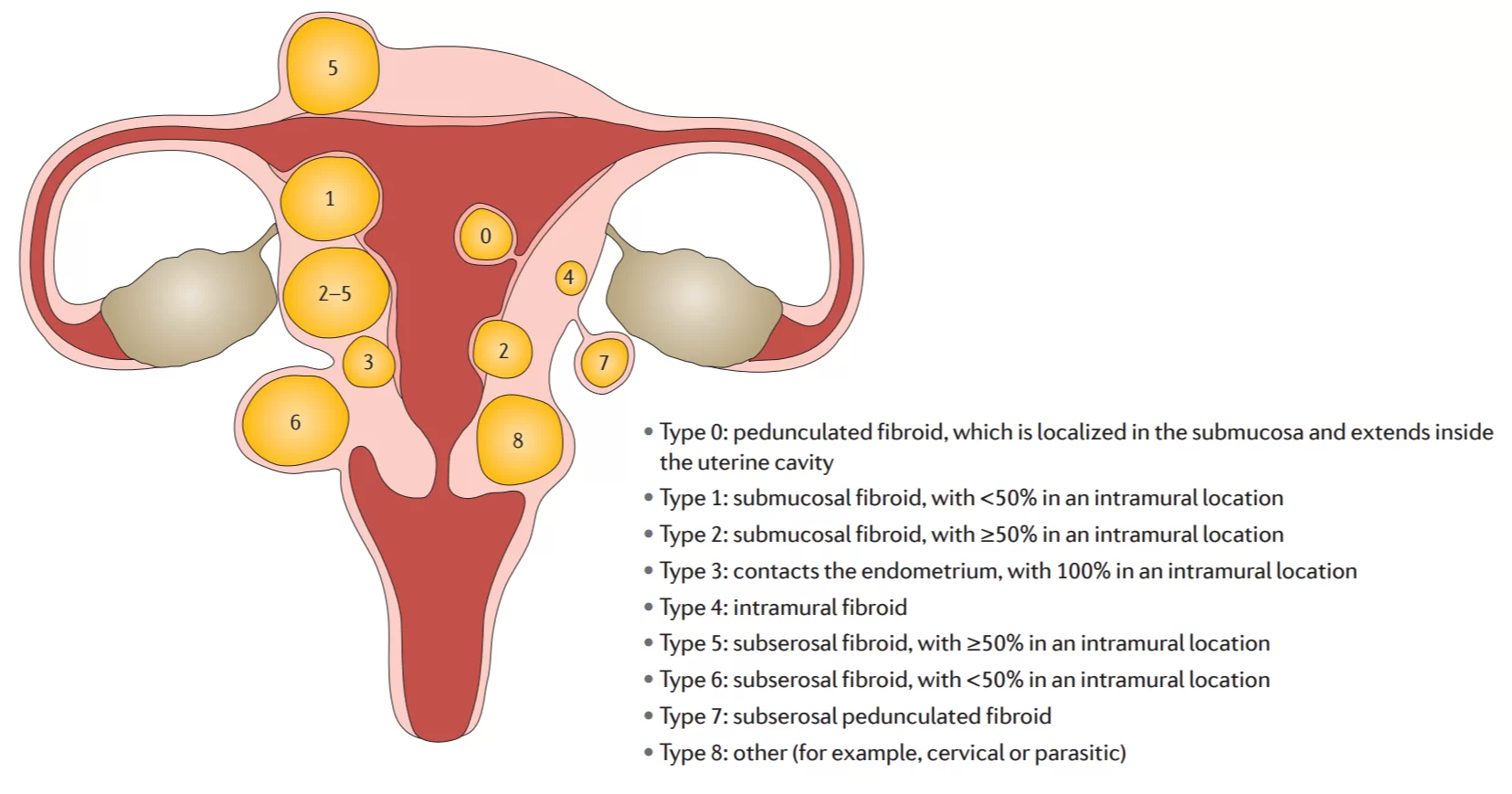
Uterine leiomyoma is the most common benign tumor in women. The prevalence of uterine leiomyoma is about 25 % in women of childbearing age and 70 % in perimenopausal women [1]. Most lesions are caused by MED12 mutations [2]. Hereditary leiomyomatosis and renal cell cancer (HLRCC) syndrome is caused by the germline mutation of fumarate hydratase ( FH ) gene, which is characterized by multiple cutaneous leiomyomas, early-onset uterine fibroids and renal cell carcinoma [3]. Because of its early characterization, easy detection and good prognosis, uterine fibroids can be used as a benign sentinel tumor for HLRCC-related renal cell carcinoma. Federation International of Gynecology and Obstetrics (FIGO) uterine leiomyoma subclassification system [4].
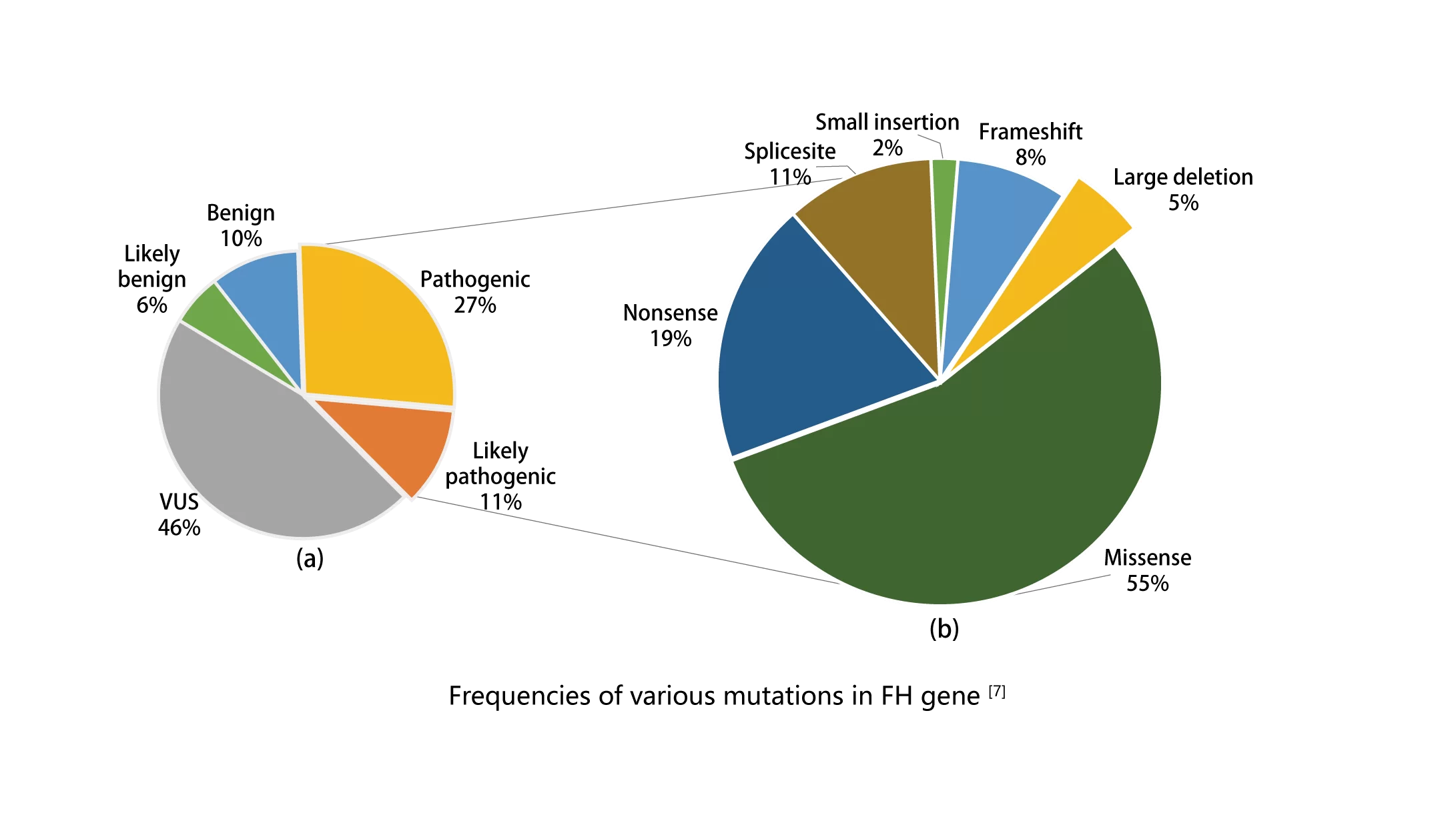
Immunohistochemistry relying solely on FH will lead to missed diagnosis in some patients, and genetic testing is the gold standard for the diagnosis of FH-deficient uterine leiomyoma [3]. The FH gene is about 22 kb in length and consists of 10 exons. Mutations have high randomness, and there is no clear hot spot of high frequency mutation [6].Copy number variation ( CNV ) is also one of the molecular genetic mechanisms of FH mutation [3]. Some FH-deficient uterine fibroids and renal cell carcinoma are caused by systemic mutations in the FH gene [3] [6]. Suspected patients need to perform whole-exome detection of germline and systemic FH mutations at the same time [6].
[1] BJOG. 2017 Sep;124(10):1501-1512.
[2] WHO Classification of Tumors of the Female Reproductive System, 5th Edition
[3] Chinese expert consensus on the diagnosis and treatment of fumarate hydratase-deficient uterine leiomyoma (2023 edition)
[4] Nat Rev Dis Primers. 2016 Jun 23;2:16043.
[5] NCCN Clinical Practice Guidelines in Kidney Cancer 2026.v1
[6] Consensus on the clinical diagnosis and treatment of fumarate hydratase-deficient renal cell carcinoma
[7] Int J Mol Sci. 2021 Jul 26;22(15):7962.
Qualitatively detect the complete coding region (CDS) of the FH gene and the MED12 gene hotspot region in tissue and/or control samples .
1、Auxiliary diagnosis of HLRCC ;
2、Suggesting the efficacy of targeted therapy in patients with FH-deficient RCC.

(1) Uterine leiomyoma patients who initially test positive for FH-related immunohistochemistry.
(2) Uterine leiomyoma patients who are young and have severe symptoms.
(3) Young patients with type II pRCC.
(4) Patients with multiple cutaneous leiomyomas and accompanying pain.
(5) Patients with a family history of Hereditary Leiomyomatosis and Renal Cell Cancer (HLRCC) or close relatives of such individuals.
01 Nucleic Acid Extraction
02 Library Preparation(3.5 hours total time)
03 Sequencing
04 Auto-data Analysis
05 Report





 闽公网安备35021202000745
闽公网安备35021202000745